
Department of Chemistry and Biochemistry
Ruth W. Molette Professor and Chair
Research Areas: Biochemistry
Office: 179 Chemistry Building
Address:
179 Chemistry Building
Auburn, AL 36849
Phone: (334) 844-6944
Fax: (334) 844-6959
Email: goodwdc@auburn.edu
Biochemistry: Structure and function of heme-dependent enzymes
Our laboratory is interested in the relationship between an enzyme’s structure and its catalytic function in biological systems. In particular, we focus on enzymes that require the organometallic cofactor heme in order to function. Heme is used by a surprisingly broad range of enzymes to accomplish an equally broad range of biologically essential tasks. For example, these enzymes are central to metabolizing foreign compounds, safely disposing of H2O2 (a toxic side product of aerobic metabolism), and mounting an effective immune response. In spite of the many and very different functions accomplished by heme-dependent enzymes, each of them relies on this organometallic molecule to accomplish the job. Clearly, the protein structure surrounding the heme group is what dictates the unique catalytic abilities of each heme-dependent enzyme.
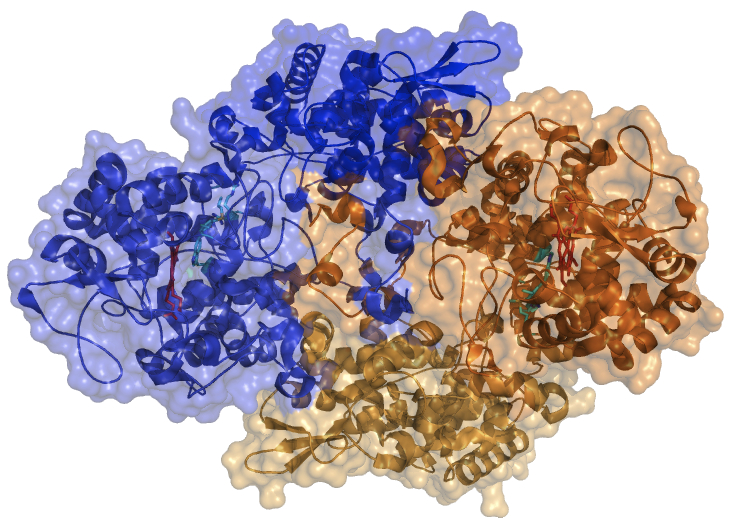 The Goodwin laboratory is using a group of bacterial enzymes called catalase-peroxidases to shed light on a poorly understood but very important aspect of the heme enzyme structure/function equation. Using these enzymes, we are demonstrating that structural components quite distant from the active site heme have a critical role in directing and fine-tuning the catalytic capabilities of heme enzymes. Not only does our research answer fundamental questions about the nature of catalysis in biological systems, but it also provides specific insight that is foundational for technological advances through enzyme engineering. The ability to engineer new enzymes for unique functions holds great promise for addressing urgent concerns that are global in their scope and impact (e.g., contamination of the environment by toxic pollutants).
The Goodwin laboratory is using a group of bacterial enzymes called catalase-peroxidases to shed light on a poorly understood but very important aspect of the heme enzyme structure/function equation. Using these enzymes, we are demonstrating that structural components quite distant from the active site heme have a critical role in directing and fine-tuning the catalytic capabilities of heme enzymes. Not only does our research answer fundamental questions about the nature of catalysis in biological systems, but it also provides specific insight that is foundational for technological advances through enzyme engineering. The ability to engineer new enzymes for unique functions holds great promise for addressing urgent concerns that are global in their scope and impact (e.g., contamination of the environment by toxic pollutants).
Our research also has implications for and applications toward substantial biomedical concerns, including antibiotic resistance and bacterial virulence. Catalase-peroxidase from Mycobacterium tuberculosis has been exploited for the activation of the front-line antitubercular agent isoniazid to its antibiotic form. Interestingly, the increasing prevelance of isoniazid resistant M. tuberculosis is strongly tied (over 70% of resistant strains) to mutations that compromise the ability of catalase-peroxidase to catalyze activation. Furthermore, a group of catalase-peroxidases have been identified as potential virulence factors in pathogens such as Escherichia coli strain O157:H7 and Yersinia pestis, both of which are recognized as high priority threats as agents of bioterrorism. Nevertheless, how these enzymes may operate as virulence factors has not been illuminated. Clearly, there are important benefits to be derived from understanding the structure and function of the catalase-peroxidases.
-
Krewall, J.R.; Minton, L.E.; Goodwin, D.C. "KatG structure and mechanism: Using protein-based oxidation to confront the threats of reactive oxygen" In Bridging Structure and Function in Mechanistic Enzymology. J.M. Miller, Ed., 2020, American Chemical Society, Washington, DC, pp. 83-120.
-
Sahrmann, P.G.; Donnan, P.H.; Merz, K.M.; Mansoorabadi, S.O.; Goodwin, D.C. "MRP.py: A parameterizer of post-translationally modified residues" J. Chem. Inf. Model. 2020, 60, 4424.
-
Simithy, J.; Fuanta, N.R.; Alturki, M.; Hobrath, J.V.; Wahba, A. E.; Pina, I.; Rath, J.; Hamann, M.T.; DeRuiter, J.; Goodwin, D.C.; Calderon, A.I."Slow-binding inhibition of Mycobacterium tuberculosis shikimate kinase by manzamine alkaloids" Biochemistry 2018, 57, 4923.
-
Simithy, J.; Fuanta, N.R.; Kochanowska-Karamayan, A.; Hobrath, J.V.; Hamann, M.T.; Goodwin, D.C.; Calderon, A.I. "Mechanism of irreversible inhibition of Mycobacterium tuberculosis shikimate kinase by ilimaquinone" Biochim. Biophys. Acta 2018, 1866, 731.
-
Alturki, M.S.; Fuanta, N.R.; Jarrard, M.A.; Hobrath, J.V.; Goodwin, D.C.; Rants'o, T.A.; Calderon, A.I. "A multifaceted approach to identify non-specific enzyme inhibition: Application to Mycobacterium tuberculosis shikimate kinase" Bioorg. Medicin. Chem. Lett. 2018, 28, 802.
-
Njuma, O.J.; Davis, I.; Ndontsa, E.N.; Krewall, J.R.; Liu, A.; Goodwin, D.C. "Mutual synergy between catalase and peroxidase activities of the bifunctional enzyme KatG is facilitated by electron-hole hopping within the enzyme" J. Biol. Chem. 2017, 292, 18408.
-
Kudalkar, S.N.; Njuma, O.J.; Li, Y.; Muldowney, M.; Fuanta, N.R.; Goodwin, D.C. "A role for catalase-peroxidase large loop 2 revealed by deletion mutagenesis: Control of active site water and ferric enzyme reactivity" Biochemistry 2015, 54, 1648.
-
Simithy, J.; Gill, G.; Wang, Y.; Goodwin, D.C.; Calderón, A.I. "Development of an ESI-LC-MS based assay for kinetic evaluation of M. tuberculosis shikimate kinase" Anal. Chem. 2015, 87, 2129.
-
Huang, J.; Smith, F., Panizzi, J.R.; Goodwin, D.C.; Panizzi, P. "Inactivation of myeloperoxidase by benzoic acid hydrazide" Arch. Biochem. Biophys. 2015, 570, 14.
-
McCarty, S.E.; Schellenberger, A.; Goodwin, D.C.; Fuanta, N.R.; Tekwani, B.L.; Calderón, A.I. "Plasmodium falciparum thioredoxin reductase (PfTrxR) and its role as a target for new antimalarial discovery" Molecules2015, 20, 11459.
-
Njuma, O.J.; Ndontsa, E.N.; Goodwin, D.C. "Catalase in peroxidase clothing: Interdependent cooperation of two cofactors in the catalytic versatility of KatG" Arch. Biochem. Biophys. 2014, 544, 27.
-
Wang, Y.; Goodwin D.C. "Integral role of the I'-helix in the function of the "inactive" C-terminal domain of catalase-peroxidase (KatG)" Biochim. Biophys. Acta 2013, 1834, 362.
-
Kudalkar, S.N.; Campbell, R.A.; Li, Y.; Varnado, C.L.; Prescott, C.; Goodwin, D.C. "Enhancing the peroxidatic activity of KatG by deletion mutagenesis" J. Inorg. Biochem. 2012, 116, 106.
-
Ndontsa, E.N.; Moore, R.L.; Goodwin, D.C. "Stimulation of KatG catalase activity by peroxidatic electron donors" Arch. Biochem. Biophys. 2012, 105, 215.
Last updated: 10/23/2023
