COSAM News Articles 2022 August Auburn plant biologists seek to better understand sunflower development and evolution with $750,000 NSF award
Auburn plant biologists seek to better understand sunflower development and evolution with $750,000 NSF award
Daniel S. Jones, assistant professor in the Department of Biological Sciences, is the recipient of a $752,045 award from the National Science Foundation (NSF) Division of Integrative Organismal Systems – Plant Genome Research Project. He is collaborating with Jennifer Mandel, associate professor, at the University of Memphis and John Burke, a Distinguished Research Professor and head of the Department of Plant Biology, at the University of Georgia. In total, the award is for $2.2 million among all three research groups for the next four years.
“The sunflower family (Asteraceae) account for ~10 percent of all flowering plants; however, this entire group is understudied at the genomic level,” said Jones. “Assembling new genomes from across this large family, along with its close relatives, will allow us to bridge an information gap that will directly impact future work with this important group.”
Currently, few resources are available to understand the genomic structure of the largest flowering plant family.
“Sunflowers are iconic, and people can easily identify this flower. But a sunflower ‘head’, called a capitulum, is not just a single flower,” Jones explained. “A sunflower capitulum is actually a composite of many small flowers (sometimes hundreds) arranged to appear and ecologically function as one large flower.”
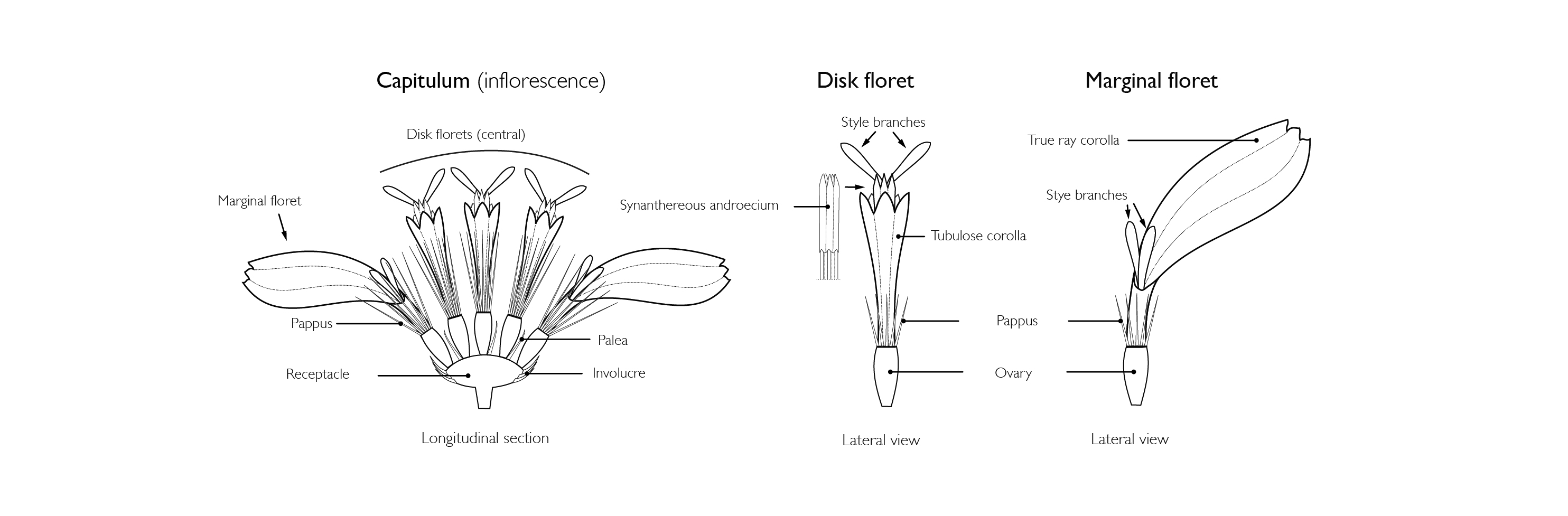
Image courtesy of Mauricio Bonifacino, collaborator from the Universidad de la República in Uruguay - https://www.compositae.org/compositae.php.
Sunflower capitula have two distinct flower types (florets).
“The outer whorl of flowers (marginal florets) are known best for their showy, long yellow ‘petals’ (called a ray). So each ray forms from a single flower” he said.
The center, where most seeds form, is where a second flower type exists.
“The center of a sunflower head is made up of disk florets; these are fairly small, but if you look closely each has five petals (ranging sometimes from yellow to dark purple) that form a tube called the corolla,” Jones added. “Each disk floret produces a single sunflower seed.”
So, each sunflower head is composed of many small flowers. This trait actually helps define the sunflower family, as plants in this group all have capitula. Commonly grown sunflower relatives that share this trait include daisies, zinnias, marigolds, coneflowers, dahlias, along with many others.
“There are approximately 32,000 species in the sunflower family (Asteraceae) and only ~300,000 described species of flowering plants,” he said.
The real-world applications of this work apply to both economic and agricultural aspects.
“Flowers are an essential aspect of many important agricultural traits as they lead directly to seeds and fruits. In this case, understanding how capitula develop can assist in efforts towards breeding larger and more structurally diverse sunflowers, impacting overall yield of seeds and seed oils,” he said.
In his lab, the CapituLab, he and his team will be working to identify and characterize different stages of flower development in sunflower relatives using fluorescent microscopy. They will then identify genes involved in capitulum development using RNA-sequencing (transcriptomics).
The CapituLab currently consists of Reid Selby (Research Assistant), Brannan Cliver and Guillian Hernández Casanova (PhD students), and Erika Lesperance and Riley Schuld (undergraduate researchers).
Jones and his team will be working on sunflower species that are grown right here on the Plains.
They grow sunflower species at the Plant Science Research Center, or PSRC, and also use growth chambers housed in the Rouse Life Sciences Building.
“The long-term goal of our lab is to develop a new genetic model system in the sunflower family to specifically help scientists understand the Asteraceae capitulum,” Jones said.
Latest Headlines
-
02/12/2025
-
02/11/2025
-
02/10/2025
-
01/30/2025
-
12/03/2024