Plants and microorganisms
rely on the shikimate pathway to produce essential aromatic
compounds. These include ubiquinone and p-aminobenzoic
acid, but most importantly, the aromatic amino acids. Because
mammals, including humans, obtain the aromatic amino acids
through the diet, they do not produce the enzymes of the
shikimate pathway, making these attractive targets for the
generation of new antitubercular agents. With our collaborator
in the Harrison School of Pharmacy, Dr.
Angela Calderon, we are generating
molecular tools to aid in the identification and
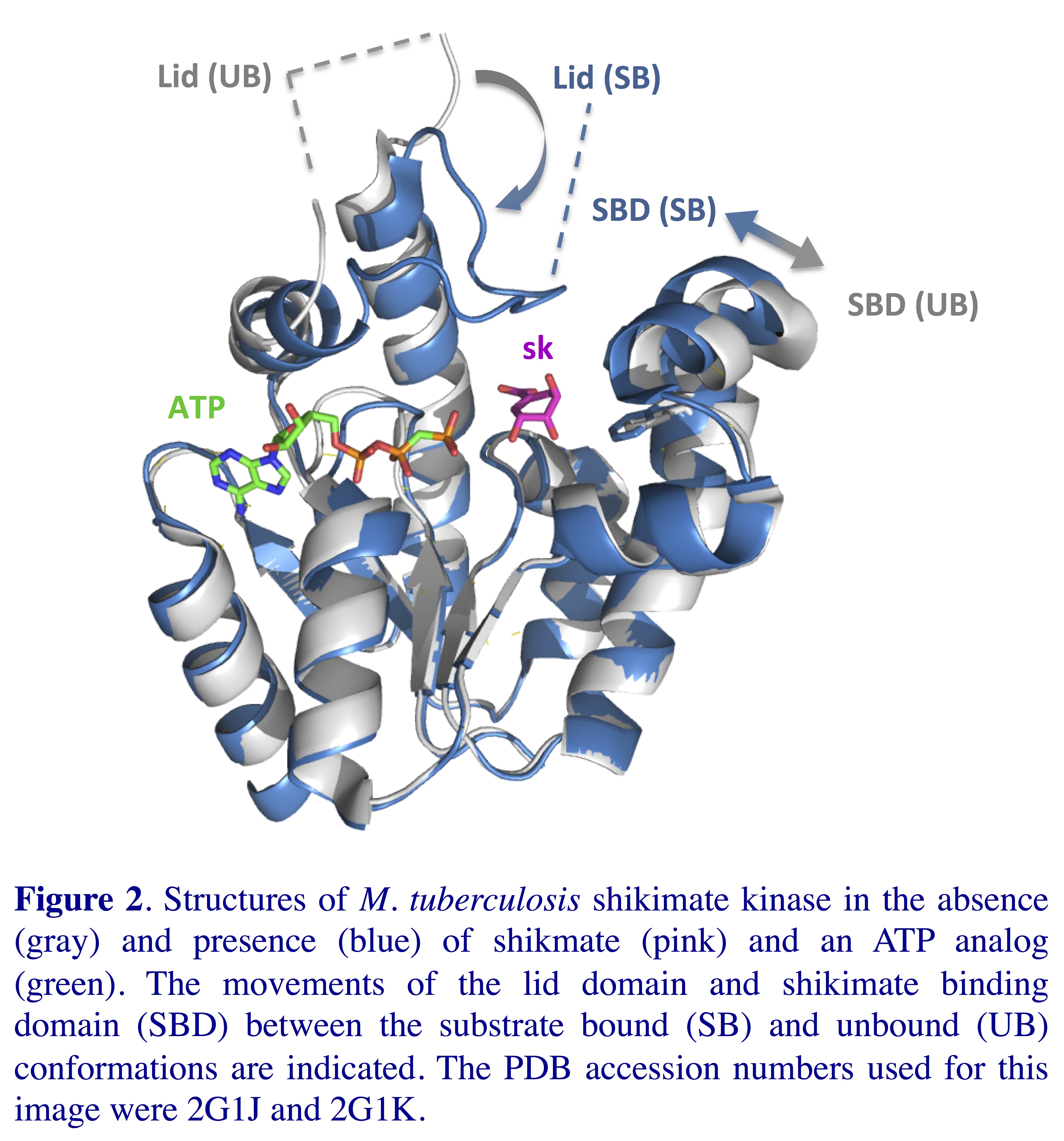
characterization of inhibitors of shikimate kinase (Figure 2),
an enzyme central to the shikimate pathway. Critical to this
endeavor, we must identify mechanistically appropriate
inhibitors. Because the enzyme operates using shikimate, a
metabolite completely absent from human metabolism, and ATP, a
metabolite found throughout human metabolism, it is important to
identify inhibitors that mimic shikimate rather than ATP in
their mechanisms of inhibition. Further, inhibitors which
exploit the known conformational dynamics of the catalytic
process are anticipated to be more effective than those that do
not. To this end, we are using mechanistically-targeted
incorporation of intrinsic protein fluorescence to produce a
panel of shikimate kinase variants that will aid in rapidly
determining the mechanism of action of candidate inhibitors.
